How to Convert JSON to IPYNB: The Complete Guide (Free & Online)
Learn how to convert JSON to IPYNB and IPYNB to JSON online for free. Covers the nbformat schema, Zeppelin migration paths, Python nbformat examples, privacy-first browser tools at jupytools.com, and when to choose JSON versus a .py export.

Quick link: run the conversion in your browser at /json-to-ipynb → (the tool). You are reading the long-form guide at /blog/how-to-convert-json-to-ipynb so the converter URL and the article URL stay distinct.
If you work in data science, machine learning, or scientific computing, you have almost certainly encountered .ipynb files. You may not know that every Jupyter Notebook is, at its core, a JSON document. This guide explains json to ipynb and ipynb to json workflows, the nbformat schema, Jupyter notebook cells and metadata, and how jupytools.com keeps sensitive notebooks on your machine while you convert—no install required.
What is an IPYNB file, and why does it look like JSON?
The .ipynb extension stands for Interactive Python Notebook. When Jupyter saves your work, it writes the entire session using the nbformat schema: structured JSON that lists every cell, output, metadata field, and kernel specification your notebook needs.
So when someone asks whether an IPYNB file is JSON—the honest answer is yes. Open any .ipynb in a plain text editor and you will see braces, arrays, and key-value pairs that follow the JSON specification. The extension is a convention so JupyterLab, VS Code, Colab, and other tools know to interpret the tree as a notebook rather than generic configuration.
Here is a minimal ipynb file example that satisfies nbformat 4:
{
"nbformat": 4,
"nbformat_minor": 5,
"metadata": {
"kernelspec": {
"display_name": "Python 3",
"language": "python",
"name": "python3"
}
},
"cells": [
{
"cell_type": "markdown",
"id": "a1b2c3d4",
"metadata": {},
"source": ["# Hello, Notebook"]
},
{
"cell_type": "code",
"execution_count": null,
"id": "e5f6g7h8",
"metadata": {},
"outputs": [],
"source": ["print('Hello, world!')"]
}
]
}
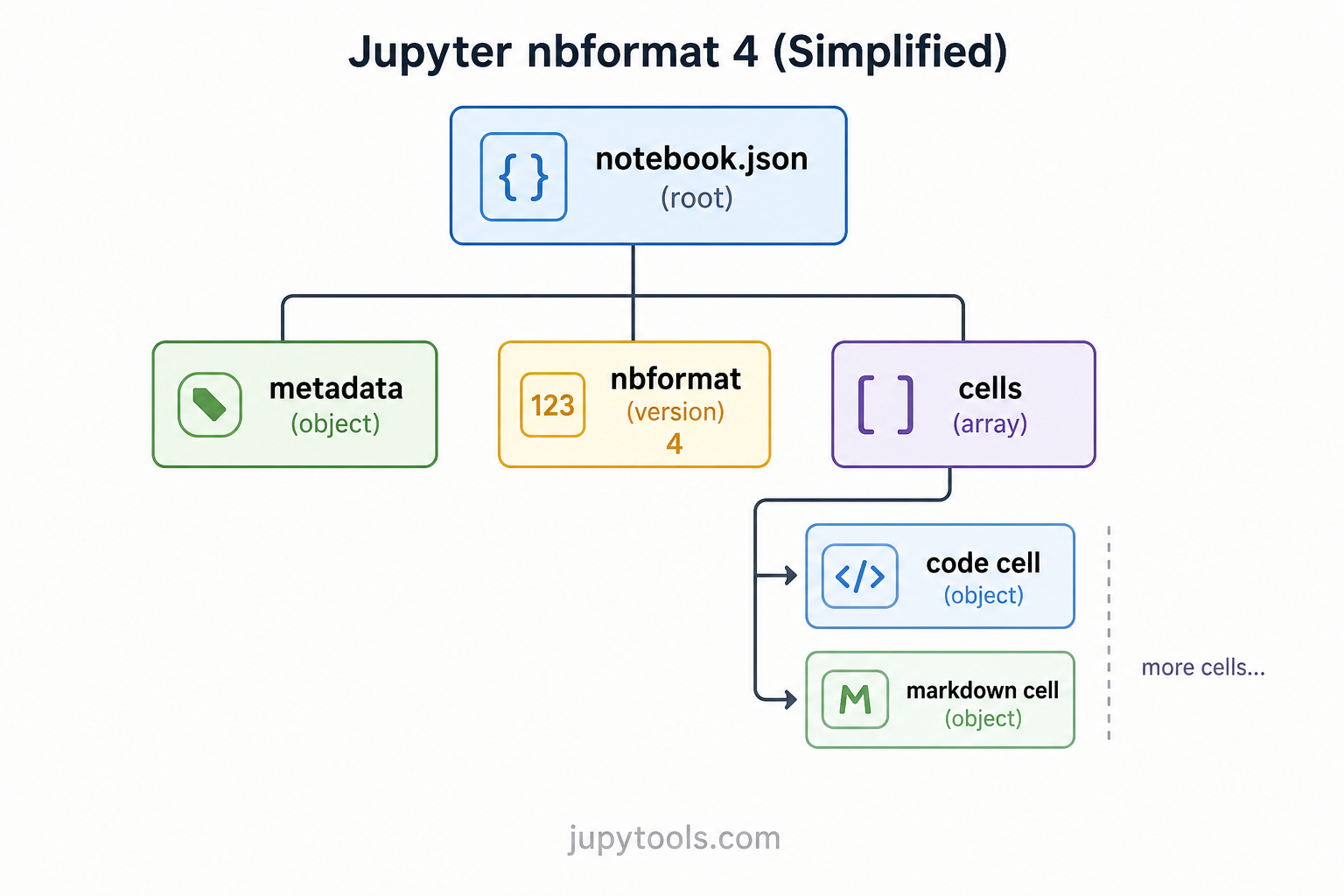
Every key has a role: nbformat pins the jupyter notebook json structure version, metadata carries kernelspec and language info, and cells is the ordered list of Markdown, code, and raw blocks. Understanding that tree is what makes convert json to ipynb predictable instead of magical.
IPYNB vs PY: what changes when you export?
Students often compare ipynb vs py when professors ask for a script, or when CI expects a single .py entrypoint.
- A
.pyfile is plain Python: statements and comments only. You run it withpython script.py. - An
.ipynbfile bundles jupyter notebook cells, Markdown narrative, saved outputs (text, HTML, images, tables), and execution metadata inside one JSON document—built for interactive exploration and notebook reproducibility.
When you convert IPYNB to Python →, most tools keep code cells as source and turn Markdown into comments; large outputs are usually dropped because a script cannot embed a live Matplotlib figure the same way a notebook cell does. When you need the full record—every stream, traceback, widget state snapshot that was saved—you export to JSON, which is exactly what ipynb to json means: the complete data science notebook format on disk.
Why convert JSON to IPYNB?
1. You have notebook JSON but the wrong extension
Sometimes a file is valid jupyter notebook json structure yet saved as .json after email gateways, LMS downloads, or API exports. The OS does not associate it with Jupyter. A dedicated json to ipynb converter validates the tree, ensures naming and structure line up with JupyterLab file format expectations, and hands you an .ipynb that opens without manual surgery.
2. You are migrating from Apache Zeppelin
Apache Zeppelin stores notebooks as Zeppelin-flavored JSON. Paragraphs, interpreters, and %md magic differ from nbformat. Converting zeppelin json to ipynb means remapping paragraphs to cells, normalizing metadata, and wrapping the result in a nbformat envelope. Community tools such as ze2nb handle the heavy mapping; once the payload matches nbformat, you can treat it like any other notebook JSON.
3. You generated a notebook programmatically
Pipelines that emit analytics reports, graded feedback, or course auto-notes often build a dict in memory, serialize to JSON, and only then need a real .ipynb that opens in jupyter notebook export tooling. Validating against nbformat before distribution prevents "works on my machine" JSON from breaking students' Jupyter installs.
How to convert JSON to IPYNB online (step by step)
The fastest json to ipynb online path—without open ipynb without jupyter turning into "install Anaconda first"—is /json-to-ipynb → on jupytools.com:
- Open the JSON to IPYNB page in Chrome, Edge, Firefox, or Safari.
- Upload your
.jsonfile (or follow on-page instructions if paste is supported). - Let the tool validate and normalize structure client-side.
- Download the
.ipynband open it in JupyterLab, classic Notebook, Colab, or VS Code.
Because processing stays inside your browser tab, confidential notebooks—API tokens, PHI in outputs, proprietary tensors—are not uploaded to a conversion farm. Pair this workflow with /ipynb-output-cleaner → when you still need to scrub cell outputs before sharing any derivative file.
How to convert IPYNB to JSON online
Engineers often need the canonical JSON to diff notebooks before merge requests, audit ipynb metadata, or feed parsers in security reviews. /ipynb-to-json → reads your .ipynb and downloads pretty-printed JSON (commonly two-space indent) so you see the real nbformat schema tree—not a flattened summary.
That JSON is also the same representation nbconvert command line workflows manipulate when you run local tooling; the browser route simply removes the Python environment requirement for one-off tasks.
Convert IPYNB to JSON using Python (nbformat)
When you need automation across dozens of files, the python nbformat library stays the standard:
import json
import nbformat
# Load the notebook
with open("my_notebook.ipynb", "r", encoding="utf-8") as f:
nb = nbformat.read(f, as_version=4)
# Write it as pretty-printed JSON
with open("my_notebook.json", "w", encoding="utf-8") as f:
json.dump(dict(nb), f, indent=2, ensure_ascii=False)
print("Done! Notebook exported as JSON.")
JSON back to IPYNB in Python
import json
import nbformat
with open("my_notebook.json", "r", encoding="utf-8") as f:
data = json.load(f)
nb = nbformat.from_dict(data)
nbformat.validate(nb)
with open("my_notebook_restored.ipynb", "w", encoding="utf-8") as f:
nbformat.write(nb, f)
print("Done! JSON restored as IPYNB.")
Calling nbformat.validate() catches missing kernelspec data, broken cell shapes, or incompatible minor versions before Jupyter surfaces a cryptic error.
Batch convert JSON files to IPYNB
import os
import json
import nbformat
os.makedirs("./output", exist_ok=True)
for filename in os.listdir("./notebooks"):
if not filename.endswith(".json"):
continue
path = os.path.join("./notebooks", filename)
with open(path, encoding="utf-8") as f:
data = json.load(f)
nb = nbformat.from_dict(data)
nbformat.validate(nb)
out_name = filename[:-5] + ".ipynb"
with open(os.path.join("./output", out_name), "w", encoding="utf-8") as f:
nbformat.write(nb, f)
Zeppelin JSON to IPYNB
Zeppelin paragraphs map conceptually to Jupyter cells, but keys, interpreters, and %md handling differ. Recommended approach:
- Install
ze2nb:pip install ze2nb. - Run
ze2nb my_zeppelin_notebook.jsonto emit.ipynb(and optional.py/.htmlcompanions). - Open the result in Jupyter and re-run cells to refresh outputs.
For tiny notebooks, manual remapping is possible—extract paragraphs, create cell_type "code" or "markdown", move text into source arrays, attach default metadata, then wrap with nbformat 4 headers—but that does not scale. Treat Zeppelin conversion as its own ETL step, then use normal notebook tooling afterward.
Copy JSON code directly into an .ipynb file
Some developers prefer raw text:
- Open the
.ipynbin VS Code, Neovim, or any UTF-8 editor. - Replace the entire file body with your JSON.
- Save with the
.ipynbextension. - Open in JupyterLab and execute cells.
This works because there is no binary blob—is ipynb a json file remains true—but invalid JSON still breaks parsing, so run nbformat.validate() or /json-to-ipynb → when you want guardrails before sharing the file.
Privacy and security when converting notebooks online
Notebooks routinely hold:
- API keys and database connection strings in code cells.
- Patient or user data inside text outputs.
- Embeddings or weights serialized in outputs.
Jupy Tools runs conversions such as IPYNB to JSON, IPYNB to PDF, IPYNB to HTML, and IPYNB to Markdown in the browser. You can confirm the privacy model by watching DevTools: no notebook payload should traverse the wire for those client-side flows. Still follow hygiene: scrub secrets before publishing any export, and use /ipynb-repair → when corruption—not confidentiality—is the issue.
Other Jupy Tools converters you might need
| Tool | Use case |
| --- | --- |
| /ipynb-to-python | Runnable .py from code cells |
| /ipynb-to-html | Self-contained browser page |
| /ipynb-to-pdf | Fixed-layout hand-ins |
| /ipynb-to-markdown | Docs portals and READMEs |
| /ipynb-to-docx | Editable Word documents |
| /ipynb-to-slides | Reveal.js style decks |
| /ipynb-merger | Combine multiple notebooks |
| /ipynb-splitter | Split on headings |
| /ipynb-diff | Side-by-side notebook review |
| /ipynb-compressor | Shrink before Git pushes |
Browse the full catalog from /tools → whenever you outgrow a single format.
Frequently asked questions
Is an IPYNB file the same as a JSON file?
Technically yes: .ipynb bytes are JSON governed by nbformat. Semantically, the extension tells editors to render jupyter notebook cells, outputs, and metadata instead of treating the file like a config blob.
How do I convert JSON to IPYNB without installing Python?
Use jupytools.com/json-to-ipynb. Upload valid notebook JSON, download .ipynb, and open it in your preferred Jupyter front end.
How do I convert IPYNB to JSON from the terminal?
Either a one-liner with python -c "import nbformat, json; ..." or a small script as shown earlier. For a GUI path, /ipynb-to-json mirrors the output without maintaining virtualenvs on every laptop.
Can I rename .json to .ipynb without conversion?
Only if the document already satisfies nbformat. Arbitrary JSON from other apps still needs transformation.
Does IPYNB to JSON keep images?
Yes—image outputs typically live as base64 payloads inside outputs arrays. That is why JSON is heavier but faithful compared with /ipynb-to-python.
Conclusion
The .ipynb format is JSON—always has been. Whether you are restoring a mis-labeled file, migrating Zeppelin notebooks through ze2nb, preparing Git diffs, or validating programmatically generated structures, the recipe is stable: understand nbformat, map fields correctly, validate, then let Jupy Tools on jupytools.com handle json to ipynb online work in the browser so you spend energy on science, not on format archaeology.
Ready to convert? Start with https://jupytools.com/json-to-ipynb for JSON → IPYNB and https://jupytools.com/ipynb-to-json for the reverse direction—free, private, and built for real notebook workflows.
Frequently asked questions
FAQ: JSON to IPYNB and IPYNB to JSON
Short answers grounded in how browser-based conversion works. Open any item to jump to detail.

